Definition
The Fréchet distance is usually explained using the analogy of a man walking his dog.
A man is walking a dog on a leash: the man can move on one curve, the dog on the other; both may vary their speed, but backtracking is not allowed. What is the length of the shortest leash that is sufficient for traversing both curves?
Intuition
We can formally define it as:
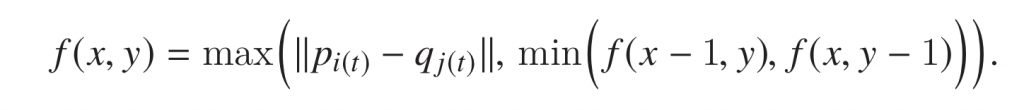
Given two sequence of points in \(\mathbb{R}^d \), p={p1, p2, p3…..pn} and q= {q1, q2, q3…. qm}, Fréchet distance represented by \(f(x,y) \) is the maximum of the minimum distances between points pi and qi. To understand it more intuitively, let us look at a simple example.
In the below graph, I have two set of polylines with positions:
\(P = {(2,1) , (3,1), (4,2), (5,1)} \)
\(Q = {(2,0),(3,0),(4,0)} \)
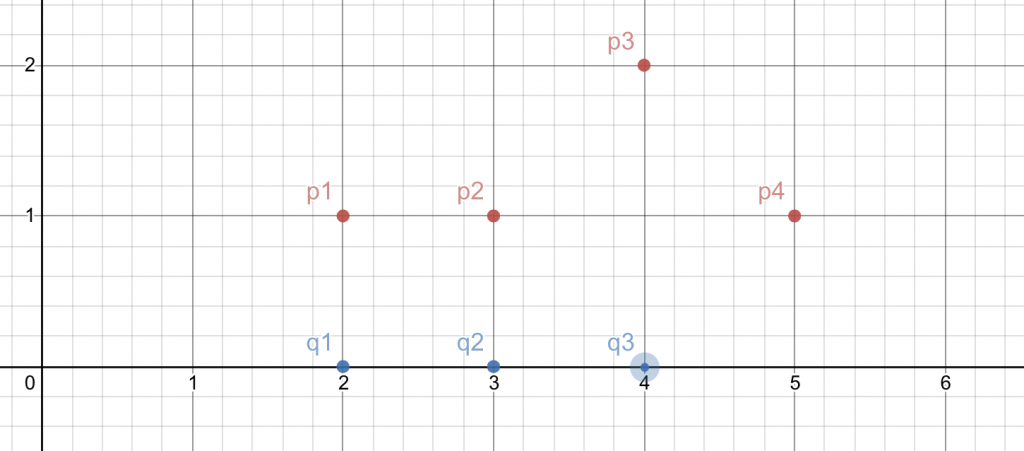
Starting and ending points being (p1,q1) and (p4,q3) respectively.
We find all the pairs of walks between p and q given as,
\(p1 -> (p1, q1) , (p1,q2), (p1,q3)\)
\(p2 -> (p2, q1), (p2,q2), (p2,q3) \)
\(p3 -> (p3, q2), (p3,q2), (p3,q3) \)
\(p4 -> (p4, q2), (p4,q2), (p4,q3) \)
then, we find the minimum distances for each pair.
\(min\{p1\} = min\{dist(p1,q1), dist(p1,q2), dist(p1,q3)\} = min\{1, 1.4, 2.2\} = 1 \)
\(min\{p2\} = min\{dist(p2,q1), dist(p2,q2), dist(p2,q3)\} = min\{1.4, 1, 1.4\} = 1\)
\(min\{p3\} = min\{dist(p3,q1), dist(p3,q2), dist(p3,q3)\} = min\{2.8, 2.2, 2\} = 2 \}\)
\(min\{p4\} = min\{dist(p3,q1), dist(p3,q2), dist(p3,q3)\} = min\{3.2, 2.2, 2\} = 2\)
Now, we can find the Fréchet distance by finding the maximum of all the minimums we calculated in the last step.
\(f(x,y) = max\{min\{p1\}, min\{p2\}, min\{p3\}, min\{p4\}\} = max\{1,1,2,2\} = 2 \)
Implementation
The implementation described by Thomas Eiter and Heikki Mannila [1] uses dynamic programming to reduce the time complexity of finding the Fréchet distance by navigating through only three possible paths.
- P and Q both move one step
- P stays where it is, Q moves one step
- P moves one step, Q stays where it is
This reduces the total number of pairs for each point in P. The implementation is as below:
from scipy.spatial.distance import euclidean
import seaborn as sns
P = [[2,1] , [3,1], [4,2], [5,1]]
Q = [[2,0] , [3,0], [4,0]]
p_length = len(P)
q_length = len(Q)
distance_matrix = np.ones((p_length, q_length)) * -1
# fill the first value with the distance between
# the first two points in P and Q
distance_matrix[0, 0] = euclidean(P[0], Q[0])
# load the first column and first row with distances (memorize)
for i in range(1, p_length):
distance_matrix[i, 0] = max(distance_matrix[i-1, 0], euclidean(P[i], Q[0]))
for j in range(1, q_length):
distance_matrix[0, j] = max(distance_matrix[0, j-1], euclidean(P[0], Q[j]))
for i in range(1, p_length):
for j in range(1, q_length):
distance_matrix[i, j] = max(min(distance_matrix[i-1, j], distance_matrix[i, j-1], distance_matrix[i-1, j-1]),
euclidean(P[i], Q[j]))
distance_matrix[p_length-1, q_length-1]
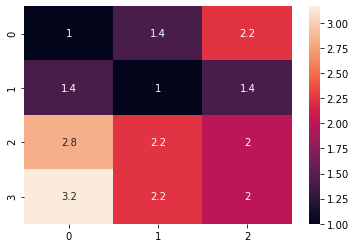
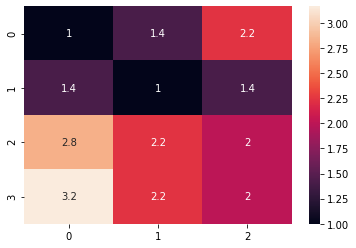
sns.heatmap(distance_matrix, annot=True)The preview of our sample as a heatmap is as below:
References:
[1] Thomas Eiter and Heikki Mannila. Computing discrete Frechet
distance. Technical report, 1994. http://www.kr.tuwien.ac.at/staff/eiter/et-archive/cdtr9464.pdf
[2] Jupyter Notebook with Frechet distance implementation.
https://colab.research.google.com/drive/1Cht2BSmKs6vCNfQZisoxMFZ_9eYMDrIi